PyEGRO.sensitivity.SApce Usage Examples¶
This document provides examples of how to use the PyEGRO.sensitivity.SApce module for sensitivity analysis using Polynomial Chaos Expansion (PCE).
Table of Contents¶
- Quick Start
- Basic Setup
- Running the Analysis
- Working with Different Models
- Using a Direct Function
- Using a Surrogate Model
- Example Applications
- Ishigami Function
- Rosenbrock Function
- Borehole Function
- Circuit Design Problem
Quick Start¶
Basic Setup¶
The simplest way to use the PyEGRO.sensitivity.SApce module is with the convenience function run_sensitivity_analysis.
import numpy as np
from PyEGRO.sensitivity.SApce import run_sensitivity_analysis
# Define a simple test function
def test_function(X):
"""
Simple quadratic function with interactions.
"""
x1 = X[:, 0]
x2 = X[:, 1]
x3 = X[:, 2]
return x1**2 + 0.5*x2**2 + 0.1*x3**2 + 0.7*x1*x2 + 0.2*x2*x3
# Define data_info with variable definitions
data_info = {
'variables': [
{
'name': 'x1',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-1, 1],
'description': 'First input variable'
},
{
'name': 'x2',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-1, 1],
'description': 'Second input variable'
},
{
'name': 'x3',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-1, 1],
'description': 'Third input variable'
}
]
}
# Save data_info to a JSON file
import json
import os
os.makedirs("DATA_PREPARATION", exist_ok=True)
with open("DATA_PREPARATION/data_info.json", 'w') as f:
json.dump(data_info, f)
# Run the analysis
results_df = run_sensitivity_analysis(
data_info_path="DATA_PREPARATION/data_info.json",
true_func=test_function,
polynomial_order=3,
num_samples=500,
output_dir="RESULT_SA_QUADRATIC",
random_seed=42,
show_progress=True
)
# Print the results
print("\nSensitivity Indices:")
print(results_df)
Running the Analysis¶
For more control over the analysis process, you can use the PCESensitivityAnalysis class directly:
from PyEGRO.sensitivity.SApce import PCESensitivityAnalysis
# Initialize the analysis
analyzer = PCESensitivityAnalysis(
data_info_path="DATA_PREPARATION/data_info.json",
true_func=test_function,
output_dir="RESULT_SA_DETAILED",
show_variables_info=True,
random_seed=42
)
# Run the analysis
results_df = analyzer.run_analysis(
polynomial_order=4, # Higher order for more accuracy
num_samples=1000,
show_progress=True
)
# Examine the summary statistics
import json
with open("RESULT_SA_DETAILED/analysis_summary.json", 'r') as f:
summary = json.load(f)
print("\nAnalysis Summary:")
for key, value in summary.items():
print(f"{key}: {value}")
Working with Different Models¶
Using a Direct Function¶
For analytical functions or inexpensive simulations, you can directly use the function for evaluations:
import numpy as np
from PyEGRO.sensitivity.SApce import run_sensitivity_analysis
# Define a complex function with interactions
def complex_function(X):
"""
Function with various non-linear effects and interactions.
"""
x1, x2, x3, x4 = X[:, 0], X[:, 1], X[:, 2], X[:, 3]
# Different term types
linear = 2*x1 + x2
quadratic = 3*x1**2 + 0.5*x2**2 + 0.1*x3**2
interaction = 2*x1*x2 + 0.7*x2*x3 + 0.3*x3*x4
non_linear = np.sin(x1) + np.exp(0.1*x2) + np.sqrt(np.abs(x3)) + np.log(np.abs(x4) + 1)
return linear + quadratic + interaction + non_linear
# Define data_info
data_info = {
'variables': [
{
'name': 'x1',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-1, 1],
'description': 'First variable'
},
{
'name': 'x2',
'vars_type': 'design_vars',
'distribution': 'normal',
'range_bounds': [-2, 2],
'cov': 0.1,
'description': 'Second variable'
},
{
'name': 'x3',
'vars_type': 'env_vars',
'distribution': 'normal',
'mean': 0,
'std': 1,
'description': 'Third variable'
},
{
'name': 'x4',
'vars_type': 'env_vars',
'distribution': 'uniform',
'low': -2,
'high': 2,
'description': 'Fourth variable'
}
]
}
# Save data_info
import json
with open("DATA_PREPARATION/complex_data_info.json", 'w') as f:
json.dump(data_info, f)
# Run analysis
results_df = run_sensitivity_analysis(
data_info_path="DATA_PREPARATION/complex_data_info.json",
true_func=complex_function,
polynomial_order=4,
num_samples=1000,
output_dir="RESULT_SA_COMPLEX",
show_progress=True
)
# Print top influential variables
sorted_results = results_df.sort_values('Total-order', ascending=False)
print("\nVariables Ranked by Total-order Influence:")
print(sorted_results[['Parameter', 'Total-order', 'First-order']])
Using a Surrogate Model¶
For computationally expensive models, you can use a surrogate model:
import numpy as np
from PyEGRO.sensitivity.SApce import run_sensitivity_analysis
from PyEGRO.meta.gpr.gpr_utils import DeviceAgnosticGPR
# Load a pre-trained surrogate model
model_handler = DeviceAgnosticGPR(prefer_gpu=True)
model_handler.load_model('RESULT_MODEL_GPR')
# Run analysis with the surrogate model
results_df = run_sensitivity_analysis(
data_info_path="DATA_PREPARATION/data_info.json",
model_handler=model_handler,
polynomial_order=5, # Can use higher order since evaluations are cheap
num_samples=2000, # Can use more samples since evaluations are cheap
output_dir="RESULT_SA_SURROGATE",
show_progress=True
)
# Examine first-order vs total-order differences to detect interactions
interaction_strength = results_df['Total-order'] - results_df['First-order']
results_df['Interaction_Strength'] = interaction_strength
print("\nVariable Interactions:")
for i, row in results_df.iterrows():
print(f"{row['Parameter']}: Interaction Strength = {row['Interaction_Strength']:.4f}")
Example Applications¶
Ishigami Function¶
The Ishigami function is a common benchmark for sensitivity analysis:
import numpy as np
from PyEGRO.sensitivity.SApce import run_sensitivity_analysis
# Define the Ishigami function
def ishigami_function(X):
"""
Ishigami function - common benchmark for sensitivity analysis.
"""
a = 7
b = 0.1
x1 = X[:, 0]
x2 = X[:, 1]
x3 = X[:, 2]
return np.sin(x1) + a * np.sin(x2)**2 + b * x3**4 * np.sin(x1)
# Define data_info
ishigami_data = {
'variables': [
{
'name': 'x1',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-np.pi, np.pi],
'description': 'First variable'
},
{
'name': 'x2',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-np.pi, np.pi],
'description': 'Second variable'
},
{
'name': 'x3',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-np.pi, np.pi],
'description': 'Third variable'
}
]
}
# Save data_info
import json
import os
os.makedirs("DATA_PREPARATION", exist_ok=True)
with open("DATA_PREPARATION/ishigami_data.json", 'w') as f:
json.dump(ishigami_data, f)
# Run analysis
results_df = run_sensitivity_analysis(
data_info_path="DATA_PREPARATION/ishigami_data.json",
true_func=ishigami_function,
polynomial_order=6, # Higher order needed for trigonometric functions
num_samples=1000,
output_dir="RESULT_SA_ISHIGAMI",
show_progress=True
)
# Print results
print("\nIshigami Function Sensitivity Analysis Results:")
print("-" * 50)
print(results_df)
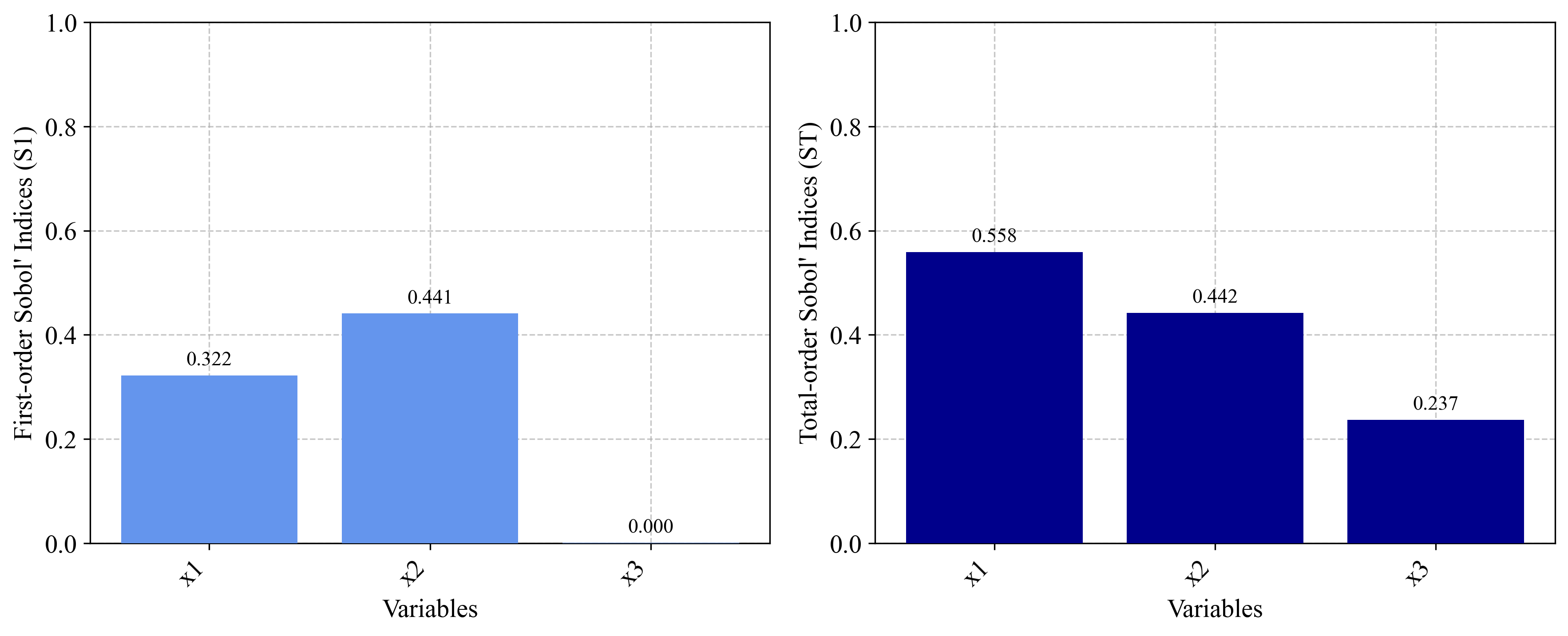
# Validate against known analytical values
print("\nAnalytical Comparison:")
print("First-order indices should be approximately:")
print("x1: 0.314, x2: 0.442, x3: 0.0")
print("Total-order indices should be approximately:")
print("x1: 0.558, x2: 0.442, x3: 0.244")
Rosenbrock Function¶
The Rosenbrock function demonstrates sensitivity analysis on an optimization benchmark:
import numpy as np
from PyEGRO.sensitivity.SApce import run_sensitivity_analysis
# Define the Rosenbrock function
def rosenbrock_function(X):
"""
Rosenbrock function - common optimization benchmark.
f(x,y) = (a-x)² + b(y-x²)²
"""
n_dim = X.shape[1]
result = 0
for i in range(n_dim - 1):
a = 1
b = 100
result += (a - X[:, i])**2 + b * (X[:, i+1] - X[:, i]**2)**2
return result
# Define data_info for 4D Rosenbrock
rosenbrock_data = {
'variables': [
{
'name': 'x1',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-2, 2],
'description': 'First dimension'
},
{
'name': 'x2',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-2, 2],
'description': 'Second dimension'
},
{
'name': 'x3',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-2, 2],
'description': 'Third dimension'
},
{
'name': 'x4',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [-2, 2],
'description': 'Fourth dimension'
}
]
}
# Save data_info
import json
with open("DATA_PREPARATION/rosenbrock_data.json", 'w') as f:
json.dump(rosenbrock_data, f)
# Run analysis
results_df = run_sensitivity_analysis(
data_info_path="DATA_PREPARATION/rosenbrock_data.json",
true_func=rosenbrock_function,
polynomial_order=5,
num_samples=1000,
output_dir="RESULT_SA_ROSENBROCK",
show_progress=True
)
# Print results
print("\nRosenbrock Function Sensitivity Analysis Results:")
print("-" * 50)
print(results_df)
# Calculate interaction effects
interaction_effect = results_df['Total-order'] - results_df['First-order']
results_df['Interaction_Effect'] = interaction_effect
# Print interaction analysis
print("\nVariable Interaction Analysis:")
for i, row in results_df.iterrows():
if row['Interaction_Effect'] > 0.1:
print(f"{row['Parameter']}: Strong interaction effect ({row['Interaction_Effect']:.4f})")
elif row['Interaction_Effect'] > 0.01:
print(f"{row['Parameter']}: Moderate interaction effect ({row['Interaction_Effect']:.4f})")
else:
print(f"{row['Parameter']}: Weak interaction effect ({row['Interaction_Effect']:.4f})")
Borehole Function¶
The borehole function is a common benchmark in engineering sensitivity analysis:
import numpy as np
from PyEGRO.sensitivity.SApce import run_sensitivity_analysis
# Define the borehole function
def borehole_function(X):
"""
Borehole function - models water flow through a borehole.
"""
rw = X[:, 0] # radius of borehole (m)
r = X[:, 1] # radius of influence (m)
Tu = X[:, 2] # transmissivity of upper aquifer (m²/year)
Hu = X[:, 3] # potentiometric head of upper aquifer (m)
Tl = X[:, 4] # transmissivity of lower aquifer (m²/year)
Hl = X[:, 5] # potentiometric head of lower aquifer (m)
L = X[:, 6] # length of borehole (m)
Kw = X[:, 7] # hydraulic conductivity of borehole (m/year)
numerator = 2 * np.pi * Tu * (Hu - Hl)
denominator = np.log(r / rw) * (1 + (2 * L * Tu) / (np.log(r / rw) * rw**2 * Kw) + Tu/Tl)
return numerator / denominator
# Define data_info
borehole_data = {
'variables': [
{
'name': 'rw',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [0.05, 0.15],
'description': 'Radius of borehole (m)'
},
{
'name': 'r',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [100, 50000],
'description': 'Radius of influence (m)'
},
{
'name': 'Tu',
'vars_type': 'env_vars',
'distribution': 'uniform',
'low': 63070,
'high': 115600,
'description': 'Transmissivity of upper aquifer (m²/year)'
},
{
'name': 'Hu',
'vars_type': 'env_vars',
'distribution': 'uniform',
'low': 990,
'high': 1110,
'description': 'Potentiometric head of upper aquifer (m)'
},
{
'name': 'Tl',
'vars_type': 'env_vars',
'distribution': 'uniform',
'low': 63.1,
'high': 116,
'description': 'Transmissivity of lower aquifer (m²/year)'
},
{
'name': 'Hl',
'vars_type': 'env_vars',
'distribution': 'uniform',
'low': 700,
'high': 820,
'description': 'Potentiometric head of lower aquifer (m)'
},
{
'name': 'L',
'vars_type': 'design_vars',
'distribution': 'uniform',
'range_bounds': [1120, 1680],
'description': 'Length of borehole (m)'
},
{
'name': 'Kw',
'vars_type': 'env_vars',
'distribution': 'uniform',
'low': 9855,
'high': 12045,
'description': 'Hydraulic conductivity of borehole (m/year)'
}
]
}
# Save data_info
import json
with open("DATA_PREPARATION/borehole_data.json", 'w') as f:
json.dump(borehole_data, f)
# Run analysis
results_df = run_sensitivity_analysis(
data_info_path="DATA_PREPARATION/borehole_data.json",
true_func=borehole_function,
polynomial_order=3,
num_samples=1000,
output_dir="RESULT_SA_BOREHOLE",
show_progress=True
)
# Print results
print("\nBorehole Function Sensitivity Analysis Results:")
print("-" * 50)
print(results_df.sort_values('Total-order', ascending=False))
# Identify key parameters
top_params = results_df.sort_values('Total-order', ascending=False).head(3)['Parameter'].tolist()
print("\nBorehole Analysis Insights:")
print(f"Most influential parameters: {', '.join(top_params)}")
print("\nEngineering Recommendations:")
for param in top_params:
print(f"- Focus on accurate measurement and control of {param}")
Circuit Design Problem¶
This example demonstrates sensitivity analysis for an electronic circuit design:
import numpy as np
from PyEGRO.sensitivity.SApce import run_sensitivity_analysis
# Define a circuit model (RC low-pass filter)
def circuit_performance(X):
"""
Calculate the cutoff frequency and performance metrics of an RC filter.
Variables:
- R1: Resistor 1 value (ohms)
- R2: Resistor 2 value (ohms)
- C1: Capacitor 1 value (farads)
- C2: Capacitor 2 value (farads)
- f: Frequency of interest (Hz)
"""
R1 = X[:, 0]
R2 = X[:, 1]
C1 = X[:, 2]
C2 = X[:, 3]
f = X[:, 4]
# Impedances
Z_R1 = R1
Z_R2 = R2
Z_C1 = 1 / (2j * np.pi * f * C1)
Z_C2 = 1 / (2j * np.pi * f * C2)
# Parallel combination of R2 and C2
Z_parallel = (Z_R2 * Z_C2) / (Z_R2 + Z_C2)
# Series with R1 and C1
Z_total = Z_R1 + Z_C1 + Z_parallel
# Calculate transfer function magnitude
H = Z_parallel / Z_total
gain_dB = 20 * np.log10(np.abs(H))
# For sensitivity analysis, use the absolute gain loss
return -gain_dB
# Define data_info
circuit_data = {
'variables': [
{
'name': 'R1',
'vars_type': 'design_vars',
'distribution': 'normal',
'range_bounds': [980, 1020], # 1kΩ with 2% tolerance
'cov': 0.02,
'description': 'Resistor 1 (ohms)'
},
{
'name': 'R2',
'vars_type': 'design_vars',
'distribution': 'normal',
'range_bounds': [9800, 10200], # 10kΩ with 2% tolerance
'cov': 0.02,
'description': 'Resistor 2 (ohms)'
},
{
'name': 'C1',
'vars_type': 'design_vars',
'distribution': 'normal',
'range_bounds': [9.5e-9, 10.5e-9], # 10nF with 5% tolerance
'cov': 0.05,
'description': 'Capacitor 1 (farads)'
},
{
'name': 'C2',
'vars_type': 'design_vars',
'distribution': 'normal',
'range_bounds': [9.5e-9, 10.5e-9], # 10nF with 5% tolerance
'cov': 0.05,
'description': 'Capacitor 2 (farads)'
},
{
'name': 'f',
'vars_type': 'env_vars',
'distribution': 'uniform',
'low': 1000,
'high': 10000,
'description': 'Frequency (Hz)'
}
]
}
# Save data_info
import json
with open("DATA_PREPARATION/circuit_data.json", 'w') as f:
json.dump(circuit_data, f)
# Run analysis
results_df = run_sensitivity_analysis(
data_info_path="DATA_PREPARATION/circuit_data.json",
true_func=circuit_performance,
polynomial_order=4,
num_samples=800,
output_dir="RESULT_SA_CIRCUIT",
show_progress=True
)
# Print results
print("\nCircuit Design Sensitivity Analysis Results:")
print("-" * 50)
print(results_df.sort_values('Total-order', ascending=False))
# Electronics design insights
top_params = results_df.sort_values('Total-order', ascending=False).head(2)['Parameter'].tolist()
print("\nCircuit Design Insights:")
print(f"Most influential parameters: {', '.join(top_params)}")
print("\nDesign Recommendations:")
for param in top_params:
if param in ['R1', 'R2']:
print(f"- Use precision resistors for {param} with tighter tolerance")
elif param in ['C1', 'C2']:
print(f"- Use high-stability capacitors for {param} (e.g., film instead of ceramic)")
elif param == 'f':
print("- Design the circuit for the specific frequency range of interest")